Research Articles
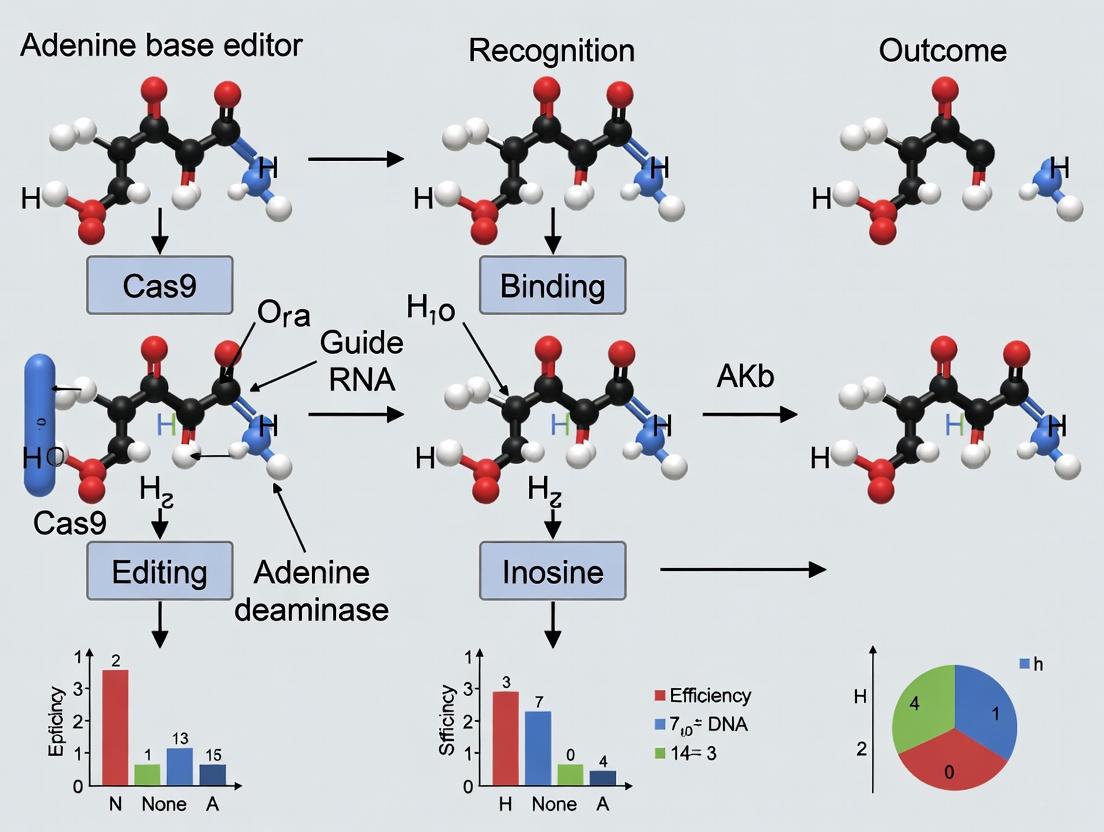
Adenine Base Editors (ABEs): A Complete Guide to Precision Genome Editing Mechanisms and Applications
This comprehensive article provides researchers, scientists, and drug development professionals with an in-depth analysis of adenine base editors (ABEs).
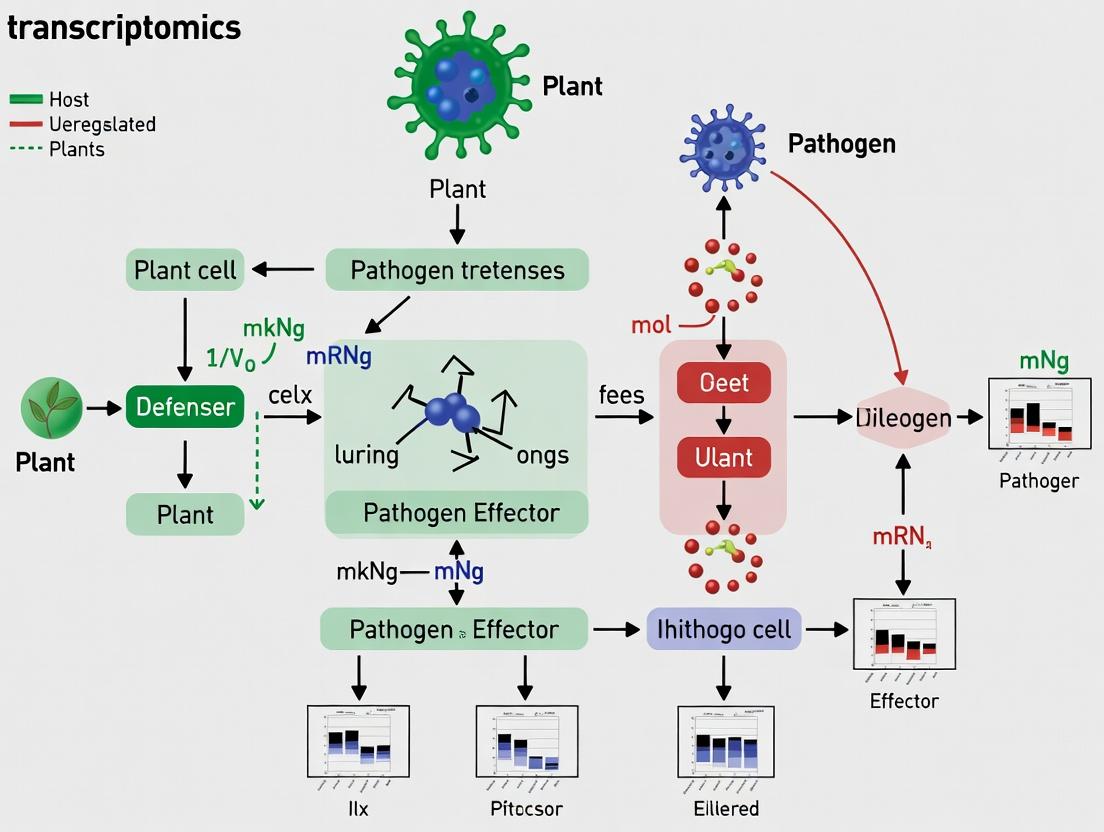
Decoding Plant Immune Dialogues: A Comprehensive Guide to Host-Pathogen Interaction Transcriptomics
This article provides a detailed exploration of transcriptomic approaches for analyzing plant-pathogen interactions, tailored for researchers and drug development professionals.
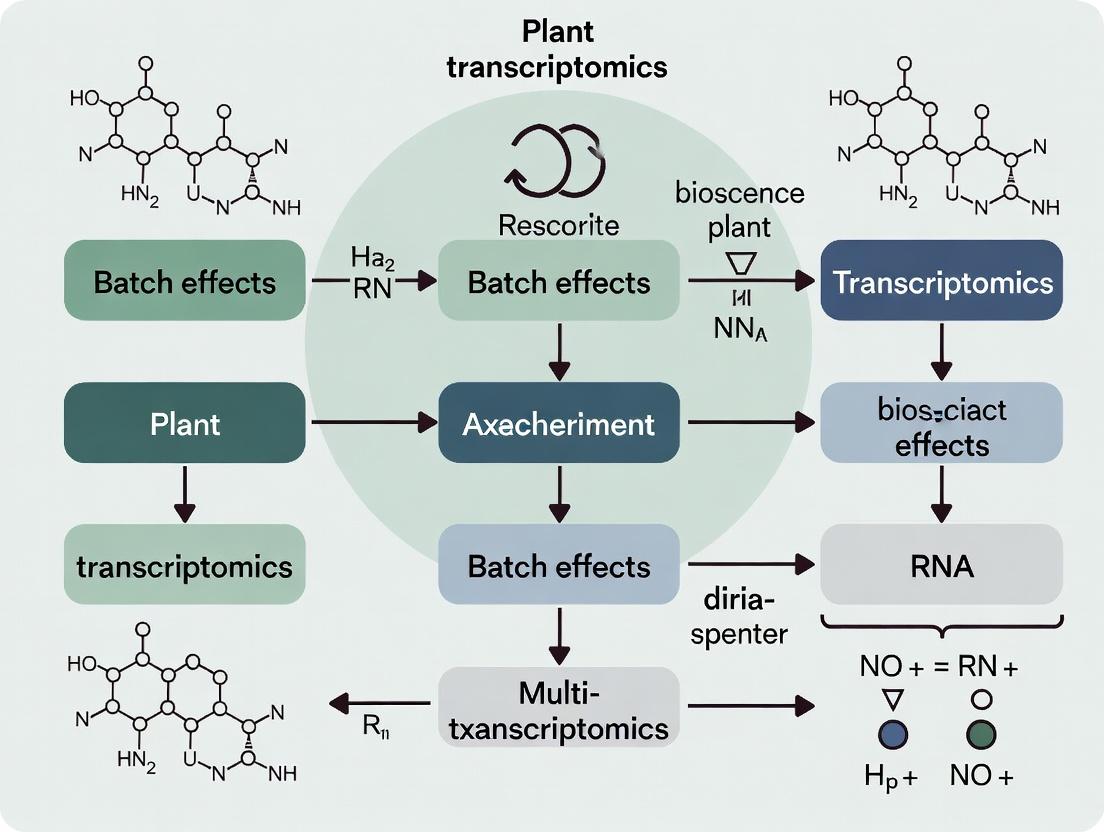
Batch Effect Correction in Plant Transcriptomics: A Comprehensive Guide for Multi-Experiment Integration
This article provides a systematic framework for identifying, diagnosing, and correcting batch effects in plant transcriptomics studies that integrate data from multiple experiments.
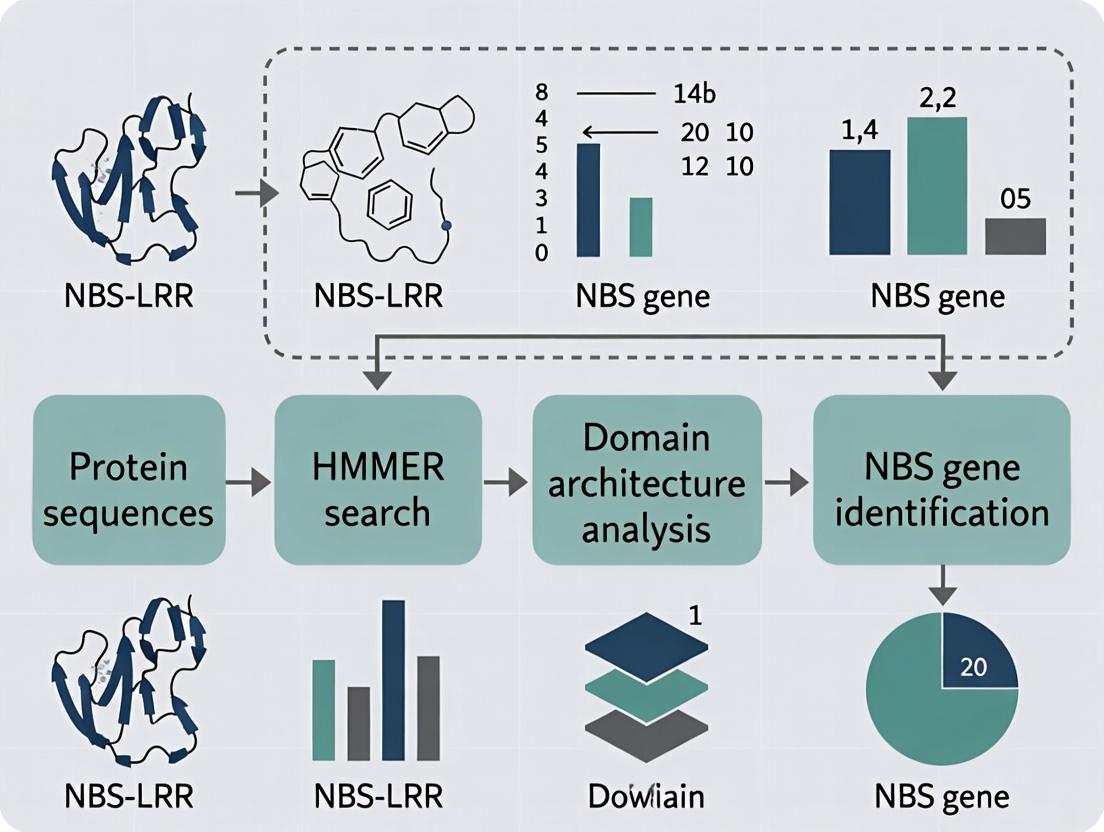
Complete HMMER Pipeline Guide: Identifying Disease-Resistant NBS Genes in Genomic Research
This comprehensive guide details the implementation of a HMMER-based bioinformatics pipeline for the accurate identification of Nucleotide-Binding Site (NBS) genes, crucial players in plant innate immunity and disease resistance.
Precision Engineering in Plants: A Comprehensive Guide to gRNA Design for CRISPR Base Editors
This article provides a systematic guide for researchers on designing effective guide RNAs (gRNAs) for plant base editing applications.
GhCLA1 and GoPGF: Dual Marker Genes for Optimizing RNAi and CRISPR Silencing Efficiency
This article provides a comprehensive guide for researchers on utilizing GhCLA1 and GoPGF as visual marker genes for rapidly assessing and optimizing gene silencing efficiency in plants.
Stable Reference Genes for VIGS: Validating GhACT7 and GhPP2A1 in Cotton Functional Genomics
This article provides a comprehensive resource for researchers utilizing Virus-Induced Gene Silencing (VIGS) in cotton (Gossypium hirsutum) and related species.
Decoding Immunity: How Gene Regulatory Networks Define Disease Resistance in Crops
This article provides a comprehensive analysis of gene regulatory networks (GRNs) that underpin disease susceptibility and resistance in crop plants.
From Expression to Regulation: A Comprehensive Guide to Gene Regulatory Network Inference in Plants Using Transcriptome Data
This article provides a systematic guide for researchers and biotech professionals on reconstructing Gene Regulatory Networks (GRNs) from plant transcriptome data.
Precision vs. Recall in GRN Inference: The Essential Guide to Evaluating Gene Regulatory Networks for Biomedical Research
This comprehensive guide provides researchers, scientists, and drug development professionals with an in-depth analysis of precision and recall metrics for evaluating Gene Regulatory Network (GRN) inference methods.